Date: Mon Dec 09 2013 - 09:12:40 MST
Hi All
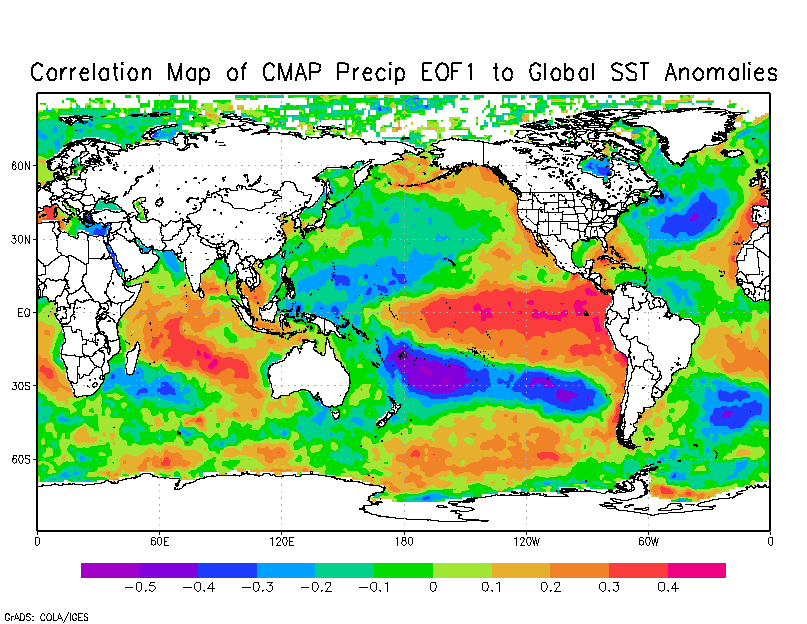
I am trying to create global maps showing the correlation of the first principle component or EOF1 of CMAP precipitation data against global SST's to pick up what area globally dominates the variability of rainfall over southern Africa. I have created a map but not sure if the coding is 100% correct as it does not look the same or similar to the map created using the climate explorer website (attached).
If someone could please have a look through my code to see if there are any major coding errors that would be most useful.
Many thanks!
Kind Regards
Melissa
Code:
; ==============================================================
; global_cor.ncl
load "$NCARG_ROOT/lib/ncarg/nclscripts/csm/gsn_code.ncl"
load "$NCARG_ROOT/lib/ncarg/nclscripts/csm/gsn_csm.ncl"
load "$NCARG_ROOT/lib/ncarg/nclscripts/csm/contributed.ncl"
begin
; ==============================================================
; User defined parameters that specify region of globe and
; ==============================================================
latS = -40.
latN = 0.
lonL = 10.
lonR = 60.
yrStrt = 1979
yrLast = 2005
season = "DJF" ; choose Dec-Jan-Feb seasonal mean
neof = 3 ; number of EOFs
optEOF = True
optEOF@jopt = 0 ; This is the default; most commonly used; no need to specify.
;;optEOF@jopt = 1 ; **only** if the correlation EOF is desired
optETS = False
; ==============================================================
; Open the file: Read only the user specified period
; ==============================================================
f = addfile ("/mnt/nfs2/geog/ml382/melphd/eof_sicz/cmap_eof.nc", "r")
lat = f->lat
TIME = f->time
YYYY = cd_calendar(TIME,-1)/100 ; entire file
iYYYY = ind(YYYY.ge.yrStrt .and. YYYY.le.yrLast)
PR = f->precip(iYYYY,:,:)
printVarSummary(PR) ; variable overview
nyrs = dimsizes(PR&time)
; =================================================================
; normalize data at each gridpoint by local standard deviation at each grid pt
; =================================================================
PR = dim_standardize_n(PR,1,0)
; =================================================================
; Reorder (lat,lon,time) the *weighted* input data
; Access the area of interest via coordinate subscripting
; =================================================================
x = PR({lat|latS:latN},{lon|lonL:lonR},time|:)
eof = eofunc_Wrap(x, neof, optEOF)
eof_ts = eofunc_ts_Wrap (x, eof, optETS)
printVarSummary( eof ) ; examine EOF variables
printVarSummary( eof_ts )
;print(eof_ts)
; ==============================================================
; Open the file: Read only the user specified period
; ==============================================================
f = addfile ("/mnt/nfs2/geog/ml382/melphd/global/oisst_globalnew.nc", "r")
sst = f->sst
;lat = f->lat
;lon = f->lon
;TIME = f->time
printVarSummary(sst) ; variable overview
sst1 =sst(:,:,:)
printVarSummary(eof_ts) ; variable overview
eof1 = eof_ts(0,:)
print(eof_ts)
; =================================================================
; Correlations calculation
; =================================================================
q = eof_ts(evn|0,time|:)
y = sst(lat|:,lon|:,time|:)
y&lat@units = "degrees_north"
y&lon@units = "degrees_east"
ccr = escorc(q, y)
printVarSummary(ccr)
ccr!0 = "lat" ; name dimensions
ccr!1 = "lon"
ccr&lat = y&lat ; assign coordinate values and
ccr&lon = y&lon ; units attributes
print(ccr)
;============================================================
; PLOTS
;============================================================
;*******************************************
; first plot
;*******************************************
wks = gsn_open_wks("X11","globalmap")
gsn_define_colormap(wks,"gui_default") ; choose colormap
rescn = True
rescn@cnFillOn = True
;rescn@gsnDraw = False ; don't draw
;rescn@gsnFrame = False ; don't advance frame
rescn@cnLevelSelectionMode = "ManualLevels" ; set manual contour levels
rescn@cnMinLevelValF = -0.4 ; set min contour level
rescn@cnMaxLevelValF = 0.4 ; set max contour level
rescn@cnLevelSpacingF = 0.2 ; set contour spacing
rescn@lbOrientation = "Vertical" ; vertical label bar
;---This resource defaults to True in NCL V6.1.0
rescn@lbLabelAutoStride = True ; optimal label stride
rescn@gsnSpreadColors = True ; use full range of colors
;rescn@gsnSpreadColorEnd = -3 ; don't use added gray
rescn@mpCenterLonF = 180. ; center plot at 180
rescn@gsnAddCyclic = True
plot = gsn_csm_contour_map(wks,ccr,rescn)
end
_______________________________________________
ncl-talk mailing list
List instructions, subscriber options, unsubscribe:
http://mailman.ucar.edu/mailman/listinfo/ncl-talk